Introduction
Gelatin methacrylate, a common photopolymerizable biomaterial, is a gelatin-based hydrogel modified with a methacrylate group that crosslinks through free radical polymerization. Derived from collagen, gelatin methacrylate exhibits many native properties of the extracellular matrix (ECM), including the presence of cell-attaching and matrix metalloproteinase responsive peptide motifs (1,2). As a photopolymerizable hydrogel with tunable mechanical properties, this material is used for a variety of purposes including 3D cellular culture. This study tests the viability of bioprinted GelMA and LAP with an Allevi bioprinter.
See Also
- Bioprinted Alginate Viability
- Bioprinted Collagen Viability
- Bioprinted PEGDA Viability
- LAP and Irgacure Viability with GelMA
- PLGA Viability Report
Results
Line structures with 400 µm thickness, 400 µm height, and spacing of 200 µm were printed with cell-laden GelMA and compared to cell-laden thin films. Live/Dead images demonstrate similar viability between thin films and bioprinted GelMA. ATP of thin films demonstrate increases in ATP from day 1 to day 7, with statistically higher ATP value when compared to day 1. The average line width was measure from Live/Dead images using ImageJ Software. After 1 day of culture, lines had an average width of 0.40 ± 0.03 mm.
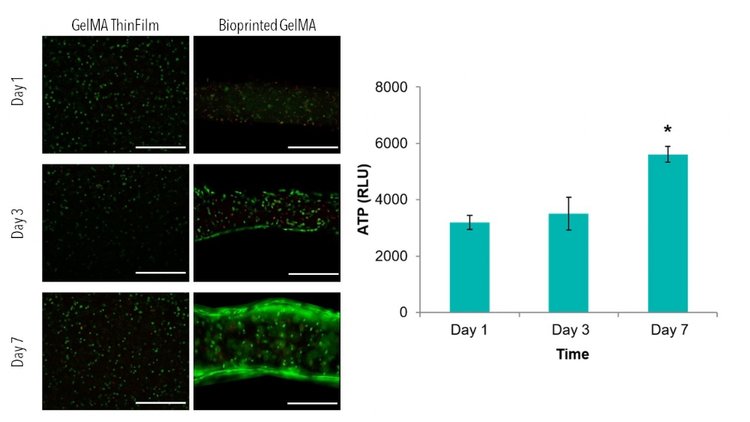
After 7 days of culture, images of bioprinted lines and thin films were taken on a confocal microscope. 3D renderings further confirmed the resolution and viability of printed structures.
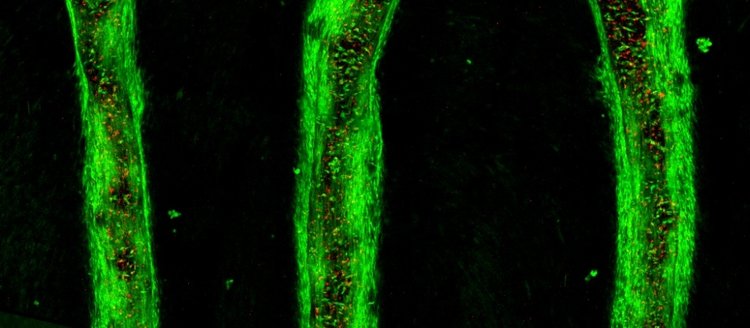
Conclusions
Allevi offers a commercially available platform with reagents, hardware, and software that allow users to create viable 3D structures with complex, reproducible geometries. Through the Allevi platform, this study demonstrates the fabrication of viable bioprinted structures composed of cell-laden GelMA and LAP.
Methods: Bioprinted GelMA & LAP
Cell Culture
Primary Human Neonatal Dermal Fibroblasts (HNDFs) obtained from ATCC were cultured in monolayer cultures at 37 °C and 5% of CO2 using Dulbecco’s Modified Eagle Medium (Corning) supplemented with 10% fetal bovine serum (Hyclone) and 1% penicillin-streptomycin-amphotericin (Corning). Passage numbers under 12 were used.
Thin Film Fabrication
To create thin films, glass coverslips were treated as suggested in the protocol provided by Allevi. Then, 0.5% (w/v) solution of LAP was dissolved in cell culture media under stirring and heat (60˚C). Next GelMA, purchased from Allevi, was mixed at 10% (w/v) in each solution. After the mixture was completely dissolved and homogenous, it was sterile filtered (0.22 μm) (Millipore) as suggested in the protocol provided by Allevi.
Once solutions were filtered, each was heated to 37°C and mixed with HNDFs at a 5 million/ml concentration. This mixture was then pipetted onto Teflon sheets (McMaster) at 10 μl. Then treated glass was placed on top of the droplets and pressed down to evenly spread the material. Next, the solution was exposed to blue light (405 nm) at 10 mW/cm². The thin films, stuck on the glass, were then removed from Teflon sheets and placed into 24 well plates (Corning).
Bioprinted Line Fabrication
First, CAD files of lines were created with solidworks. Then, these .stl files were loaded into repetier host and sliced with the print parameters suggested by Allevi, to create a .gcode print file. 2 ml of cell-laden GelMA, prepared in the same way as GelMA for fabrication of thin films, was loaded into extruder 2. 27 tapered needles were used to print structures. The lnes were printed with the sliced print file and the print parameters suggested by Allevi.
Lines were printed onto treated glass slides, then crosslinked for 2 min with blue light at 10 mW/cm². All samples were bioprinted with an Allevi 2.
Sample Analyses
To assess cell viability, CellTiter-Glo 3D ATP Assay (Promega) was performed according to manufacturer’s protocol and analyzed using a BioTek Synergy 2 Plate Reader. A LIVE/DEAD kit (Life Technologies) was used to qualitatively assess the viability of samples. Images were taken on a Nikon TE300 Inverted Fluorescent Microscope or a Leica TCS SP5 II Scanning Laser Confocal Microscope.
Statistical analysis was performed using an Analysis of Variance (ANOVA) test to determine if differences were present amongst treatment groups. If differences were determined from the ANOVA, a post hoc Tukey’s multiple comparison test was used to determine statistical differences between groups tested. A confidence level of 95% (α=0.05) was used for all analyses. Error bars on graphs represent the standard deviation from the mean.
References
- D. B. Kolesky et al, “3D Bioprinting of Vascularized, Heterogeneous Cell-laden Tissue Constructs,” Adv. Mater., vol. 26, pp. 3124-3130, 2014.
- B. D. Fairbanks et. al, “Photoinitiated Polymerization of PEG-diacrylate with lithium phenyl-2,4,6-trimethylbenzoylphosphinate: polymerization rate and cytocompatibility,” Biomaterials, vol. 30, no. 35, pp. 6702-6707, Dec 2009.

