Introduction
As a naturally occurring polysaccharide commonly derived from algae or seaweed, sodium alginate’s abundance and low cost make it a popular biomaterial for cellular encapsulation and bioprinting [1]. Sodium alginate forms a hydrogel through sodium-calcium ion exchange. As this crosslinking process occurs very quickly, sodium alginate can be difficult to bioprint on its own. Many studies combine this reagent with other materials to create a hybrid bioink or use complex multi-crosslinking steps to create 3D geometries (2-4). Here, we present a simplified process for printing sodium alginate with the use of Pluronic F127 as temporary support. However, the FRESH method of bioprinting is suggested for the best resolution and for the fabrication of more complex 3D structures. This viability report analyzes 3D structures of cell-laden bioprinted alginate with repeatable, accurate geometries.
See Also
- Bioprinted Collagen Viability
- Bioprinted GelMA and LAP Viability
- Bioprinted PEGDA Viability
- LAP and Irgacure Viability with GelMA
- PLGA Viability Report
Results
Thin Film Viability
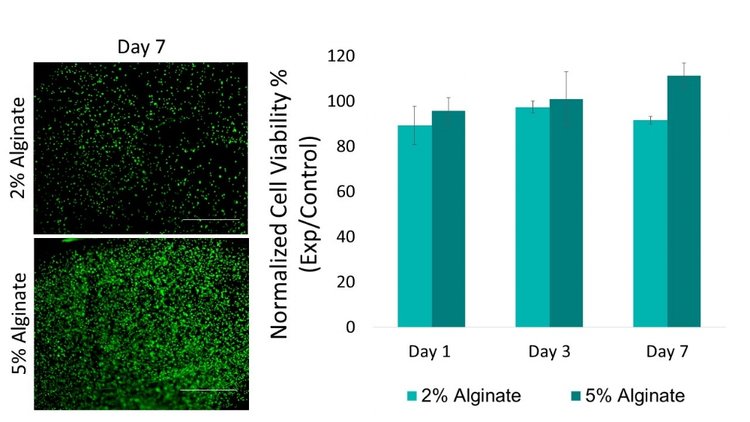
Bioprinted viability was first tested by comparing bioprinted thin films to pipetted thin films. Both 2% and 5% bioprinted thin films had over 85% viability compared to non-bioprinted controls. No statistical differences were found between 2% and 5% groups at each time point.
Bioprinted Ring Viability
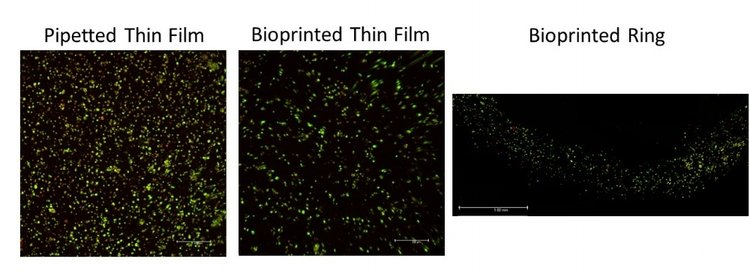
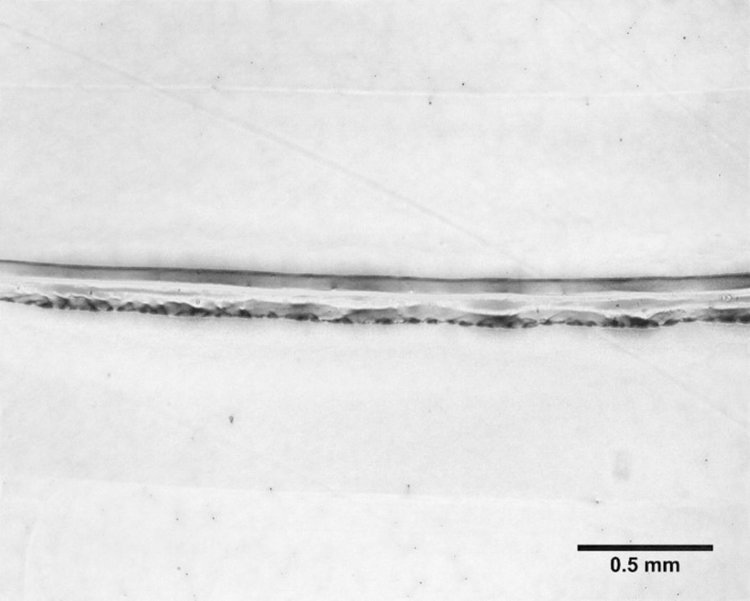
Ring structures designed with a 300 μm thickness, 400 μm height and an outer diameter of 2.3 mm were printed with 2% cell-laden alginate. The viability of these bioprinted rings were compared to bioprinted and pipetted thin films (figure 2). Qualitative analysis of live/dead imaging suggests similar viability in the bioprinted ring structures as the bioprinted and pipetted thin films. Line width averages, analyzed with ImageJ software, were 360 ± 80 μm after 1 day of culture.
Conclusions
These results demonstrate the ability of sodium alginate, calcium chloride and Pluronic F127 from Allevi to support viable cultures and create consistent, high-resolution 3D geometries when used with an Allevi 1. When used together, these reagents can create cell-laden structures with specific, repeatable 3D geometries. Further, bioprinted thin films and pipetted thin films created in this study can be used as 3D controls for future experiments with these materials.
Methods
Cell Culture
Primary Human Neonatal Dermal Fibroblasts (HNDFs) obtained from ATCC were cultured in monolayer cultures at 37 °C and 5% of CO2 using Dulbecco’s Modified Eagle Medium (Corning) supplemented with 10% fetal bovine serum (Hyclone) and 1% penicillin-streptomycin-amphotericin (Corning). Passage numbers under 12 were used.
Thin Film Fabrication
HNDFs were mixed with either 2% or 5% (w/v) alginate at a 5 million/ml concentration. Pipetted thin films were created by pipetting 10 μl of solution on transwell plates then submerging the samples in 10 mM Calcium Chloride for 10 – 15 minutes.
2% alginate bioprinted thin films and 5% alginate bioprinted thin films were printed according to the suggested print parameters. The volume of extruded bioprinted material was estimated with the volume test. Extrusion time in the printed G-code was set for extrusion of approximately 10 μl. All samples were crosslinked for 10-15 minutes in calcium chloride, then washed three times with phosphate buffered saline (PBS). All samples were bioprinted with an Allevi 2.
Bioprinted Ring Fabrication
First, CAD files of alginate rings and supporting pluronic F127 structure were created with solidworks. Then, these .stl files were loaded into repetier host and sliced with the suggested print parameters. 5 ml of Pluronic F127 was loaded into extruder 1 and 2 ml of 2% alginate with a 5 million/ml HNDF concentration was loaded into extruder 2. 30 gauge needles were used for both materials.
Rings were printed into transwell plates, then submerged in 10 mM calcium chloride first at room temperature for 5-10 minutes then on ice for another 5 minutes. After crosslinking, samples were washed with phosphate buffered saline (PBS) three times before adding media. All samples were bioprinted with an Allevi 2.
Sample Analyses
To assess cell viability, a LIVE/DEAD kit (Life Technologies) was used. Images were taken on a Nikon TE300 Inverted Fluorescent Microscope or a Leica TCS SP5 II Scanning Laser Confocal Microscope. Statistical analysis was performed using an Analysis of Variance (ANOVA) test to determine if differences were present amongst treatment groups. If differences were determined from the ANOVA, a post hoc Tukey’s multiple comparison test was used to determine statistical differences between groups tested. A confidence level of 95% (α=0.05) was used for all analyses. Error bars on graphs represent the standard deviation from the mean.
Supplementary Information
After this study, sodium alginate was tested with the FRESH method. This method was able to achieve significantly better resolution and is suggested for optimal bioprinting results with alginate.
References
- Guvendiren, Murat. Designing biomaterials for 3D printing. ACS Biomater. Sci. Eng. 2016 Article ASAP.
- Duan Bin, et al. 3D Bioprinting of Heterogeneous Aortic Valve Conduits with Alginate/Gelatin Hydrogels. J Biomed Mater Res A. 2013 May; 101(5): 1255-1264.
- Tabriz, Atabak Ghanizadeh et al. Three-dimensional bioprinting of complex cell-laden alginate hydrogel structures. Biofabrication. 2015 7 045012.
- Wu, Zhengjie et al. Bioprinting three-dimensional cell-laden tissue constructs with controllable degradation. Sci Rep. 2016 Apr 19;6:24474.

